Background and Objective: Diabetic foot ulceration is a multifactorial process involving various intrinsic complications of diabetes mellitus which cause injury to the foot at risk. The diabetic foot ulcer infections are polymicrobial in nature. Failure to recognize and control of the infectious process may have devastating consequences of limb amputation, sepsis, and mortality. Hence, the study was undertaken to determine the bacterial and clinical profile of diabetic foot ulcer using optimal culture techniques and the antimicrobial sensitivity pattern of the isolates. Materials and Methods: A total of 100 patients with diabetic foot ulcer of Wagner’s grade I and above were included in the study. Pus and tissue biopsy were collected for bacteriological study. The specimen was processed in the microbiology laboratory for Gram stain, aerobic culture, and anaerobic culture. The organisms isolated were identified by standard procedures and antimicrobial susceptibility was done by Kirby-Bauer disc diffusion method. Results: A total of 187 organisms were isolated, with an average of 1.87 organisms per specimen. Pseudomonas sp, 36 (21.9%), was the most common aerobic organisms isolated followed by Klebsiella sp, 32 (19.4%). Anaerobic organisms isolated were 22 (11.77%). The predominant anaerobic organisms isolated were Peptostreptococcus sp, 10 (45.5%). All the aerobic Gram-negative organisms were sensitive to imipenem (100%). Gram-positive organism was 100% sensitive to vancomycin. Methicillin resistant staphylococcus aureus (MRSA) was seen in 66.7%. All the anaerobes were sensitive to metronidazole, clindamycin, cefoxitin, and penicillin G . Conclusion: Pseudomonas was the most common organism isolated in our study. MRSA was seen in 66.7% of the isolate. Keywords: Aerobes, anaerobes, antibiotics, diabetic foot
Diabetic foot ulceration is a multifactorial process involving various complications of diabetes mellitus. The factors are neuropathy, abnormal foot biomechanics, and peripheral arterial disease. [1] Infection occurs following the traumatic injury with introduction of bacteria. The diabetic ulcer infections are polymicrobial, harboring anaerobic organisms synergistically with aerobes. [2] In superficial wounds, aerobic bacteria are predominant pathogens. Anaerobic organisms are found in deeper wounds. [3] Failure to recognize and control of the infectious process may have devastating consequences like limb amputation, sepsis, and mortality. [4] This study was undertaken to determine the bacterial and the clinical profile of diabetic foot ulcer.
A total of 100 patients with diabetic foot were included in the study and were classified according to Wagner’s grade. Following consent, a questionnaire was filled to record the patient’s demographic data, duration of diabetes, method of glycemic control, history of smoking, hypertension, duration of wound, and previous history of treatment. Peripheral neuropathy was assessed by touch sensation with pin prick, cotton, [5] and peripheral vascular disease by palpation of dorsalis pedis artery and posterior tibial artery. [5] Pus and tissue biopsy specimens were obtained for bacteriological study. Surface of the ulcer was rinsed with sterile normal saline, superficial exudates was debrided using a sterile instrument. Non-involved adjacent skin was sterilized with iodine and 70% alcohol. Pus was collected with the sterile cotton tipped swabs. [6] For the abscess, surface was cleansed with antiseptic and the pus was aspirated with sterile needle and syringe. Surgically, debride tissues were also obtained. Following collection, the specimen was processed in the microbiology laboratory for Gram stain, aerobic culture, and anaerobic culture. [7],[8] For aerobic culture, it was inoculated on to blood agar and MacConkey agar and incubated at 37°C for 48 hours.The organisms isolated were identified by standard biochemical reactions. For anaerobic culture, primary inoculation was done on to Robertson cooked meat medium and subcultured on blood agar and neomycin blood agar. The agar plates were incubated in anaerobic jar at 37°C for 48 hours. The organisms isolated were identified by standard procedures. [9],[10] Antibiotic susceptibility of the isolates was done by Kirby-Bauer disc diffusion method. Ampicillin (30μg), augmentin (20μg/10μg), cotrimoxazole (25μg), gentamicin (10μg), amikacin (30μg), ciprofloxacin (30μg), ceftriaxone (μg30), cefotaxime (30μg), piperacillin (100μg), imipenem (10μg), oxacillin (1μg) and vancomycin (30μg) were used. Wagner grading system for diabetic foot lesions. [11] Grade: Lesions Grade 0: No open lesions, may have deformity and cellulitis Grade 1: Superficial ulcer Grade 2: Deep ulcer to tendon, capsule, or bone Grade 3: Deep ulcer with abscess, osteomyelitis, or joint sepsis Grade 4: Localized gangrene, forefoot, or heel Grade 5: Gangrene of entire foot
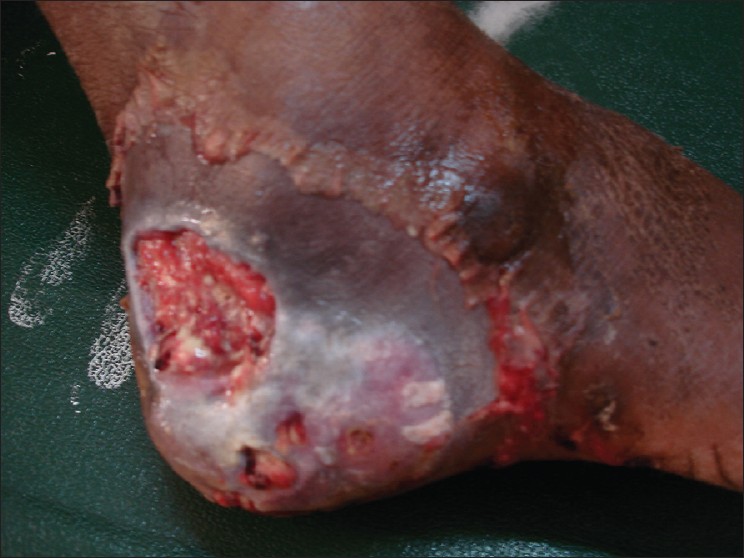
A total of 100 patients with diabetic foot of Wagner’s grade I and above were studied [Figure 1]. Among them, 65% were males and 35% were females. Most of the patients with diabetic foot were in the age group 51 to 70 (68%) years and mean age was 57.14 years SD (10.77). Fifty-seven (57%) patients were suffering from diabetes mellitus for more than five years, mean duration was 6.07 years SD (3.72). Only one (1%) case was insulin-dependent diabetes mellitus and the others were non-insulin-dependent diabetes mellitus (NIDDM) cases, 99 (99%). Thirty-nine (39%) patients presented with ulcer of 2- to 4-week duration and 30 (30%) patients presented with ulcer of 8 to 10 week duration, the mean duration of ulcer at presentation was 5.83 weeks SD (3.72). Prior to admission, 81 (81%) patients were maintained on oral hypoglycemic agents and four patients were not previously diagnosed as diabetics. Peripheral neuropathy was present in 65 (65%) and 26 (26%) had peripheral vascular disease. Sixty-three (63%) had poor glycemic control with blood sugar of more than 200 mg∕dl, five (5%) of them were hypertensive and 26 (26%) were smokers. Majority of the patient presented with the ulcer of Wagner grade II, 43 (43%), followed by grade III, 24 (24%) [Table 1].
Five patients had gangrene involving up to knee joint. One patient had gangrene of limb, chronic renal failure, and sepsis. A total of 187 organisms were isolated in the present study, with an average of 1.87 organisms per specimen. Among them, aerobic organisms isolated were 165 (88.23%) and anaerobic organisms were 22 (11.77%). Of 165 aerobic organisms isolated, Gram-negative organisms were 128 (77.6%) and Gram-positive organisms were 37 (22.4%). The most common aerobic isolates were Pseudomonas sp, 36 (21.9%), followed by Klebsiella sp, 32 (19.4%), Escherichia coli, 26 (15.8%), and Proteus sp, 23 (13.9%). Staphylococcus aureus isolated was 24 (14.5%) [Table 2].
Of 22 (11.77%) anaerobic organisms [Figure 2], the most common isolates were Peptostreptococcus sp, 10 (45.5%), followed by Bacteroides fragilis, 7 (31.8%) [Table 3].
Monomicrobial flora was present in 36 (36%) cases, of which all the isolates were aerobes. Polymicrobial flora was present in 64 (64%) cases. Of them, in 43 (43%) cases, only aerobic organisms were isolated and aerobic along with anaerobic organisms were isolated in 21 (21%) cases. In the present study, all the aerobic Gram-positive organisms were 100% sensitive to vancomycin, followed by amoxicillin/clavulanic acid (58.66%) and they were highly resistant to ampicillin (94.59%), cotrimoxazole (97.23%), and gentamicin (81.5%). MRSA was seen in 66.7% [Table 4]. Gram-negative organisms were sensitive to imipenem (100%), amikacin (57.37%), and ciprofloxacin (57.93%), except Pseudomonas was (91.7%) sensitive to imipenem and (75.8%) to piperacillin [Table 5]. All the anaerobes were sensitive to metronidazole, clindamycin, cefoxitin, and penicillin G.
Infected, nonhealing diabetic foot ulcer is the major cause for nontraumatic lower limb amputation. It is estimated to be 40 times greater than those of nondiabetics. [12] In the present study, there was significant association between diabetic foot ulceration and clinical parameters like male gender, duration of diabetes for more than 5 years, NIDDM type of diabetes mellitus, and poor control of diabetes. Other contributing factors such as neuropathy and vascular disease were present at the time of consultation. All the patients, due to lack of education on nature of illness, consulate after 2 weeks of the foot ulcer. Smoking significantly did not pose any risk for ulcer formation. Diabetic wound with multiple pathogens isolated indicate significant pathogenicity and help each other in wound degradation and early development of complication. Poor defense also leads to rapid increase in the number of microbes. Reports from various studies have shown that 39 to 90% of the diabetic foot infections are polymicrobial. [13] In the present study, polymicrobial etiology was seen in 64% of the cases. Mixed aerobes were seen in 43% cases and aerobes along with anaerobes were seen in 21% cases. Average yield per specimen was 1.87 organisms. This was nearly similar to study conducted by various workers who reported 2.2% [7] and 2.1%. [14] Pseudomonas sp, Enterococcus sp, and Proteus sp are responsible for continuing and extensive tissue destruction. [4] In the present study, Pseudomonas sp, 36 (21.9%), was the predominate isolate, followed by Klebsiella sp, 32 (19.4%), and S. aureus, 24 (14.5%). The spectrum of aerobic bacterial isolate were similar to other studies. [4],[8],[15] In contradiction, S. aureus is the most common organism in most of the study. [8],[4] In patient with Wagner’s grade I and II diabetic ulcer, S. aureus and E. coli were the common organisms isolated. In Wagner’s grade III, IV, V diabetic ulcer, four to five organisms were isolated. Pseudomonas sp, Klebsiella sp, E. coli, and S. aureus were the common isolates. Isolation and culture of anaerobic organism is tedious, complicated, requires financial resources and technology. The yield of anaerobic organisms depends on the method of sample collection and previous antibiotic intake. Various studies show that prevalence of anaerobic organisms in diabetic foot infection varies from 5% to as high as 95%. [16] With the optimal culture technique available, anaerobic organism isolated in our study was 11.77% and were associated with Wagner’s grade IV and V. The predominant anaerobic organisms isolated in our study was Peptostreptococcus sp (45.5%), followed by Bacteroides sp. Similar observations were made by various workers who reported Peptostreptococcus sp as the predominant isolate. [17],[18] A foreign study has reported Bacteroides sp as predominant isolate. [19] Treatment of diabetic foot infection requires antibiotic therapy based on the culture and surgical intervention. In our study, multiple drug resistance was observed for more than three drugs. Three Pseudomonas sp isolates were resistant to imipenem (8.3%). E. coli and Klebsiella sp showed 100% sensitivity to imipenem and nearly 70% sensitivity to ciprofloxacin, but were highly resistant to cephalosporins. An Indian study has reported 77.27% sensitivity to ciprofloxacin and 74% to cephalosporin. [8] MRSA was seen in 66.7%. This was high when compared with a South Indian study that reported 42.86%. [20] Patients with ulcer underwent local surgical debridement, five patients underwent below knee amputation and mortality was seen in one patient with gangrene of the foot, chronic renal failure, and sepsis.
Diabetic foot infection is the most common cause for hospital admission, resulting in major economic consequences for the patients, families, and society. Patient with diabetic foot ulcer requires team approach to care patient education, local care, treatment of infection with culture-guided antibiotic therapy, and surgical intervention.
Source of Support: None, Conflict of Interest: None
[Figure 1], [Figure 2]
[Table 1], [Table 2], [Table 3], [Table 4], [Table 5] |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||